Donate blood and win a hoodie!
Lend a helping hand! Donate blood between 11 March and 31 May to win a stylish hoodie.
Test if you can donate
Take a quick online test to find out whether you can donate blood or stem cells.
Take the testHealth questionnaire for blood donors
Please complete the electronic health questionnaire on the day of your appointment or, at the earliest, on the day before.
Fill out the questionnaireStem Cell Registry
A stem cell transplant is the last hope for a cure for blood cancer patients. More members are needed.
More about Stem Cell RegistryNews and articles

A Good Samaritan on a motorcycle
For decades, a Finnish motorist association, SMOTO (Suomen Motoristit ry), has been actively encouraging its members to donate blood. Raimo Ryhtä, a motorist and active blood donor from the city of Forssa,…
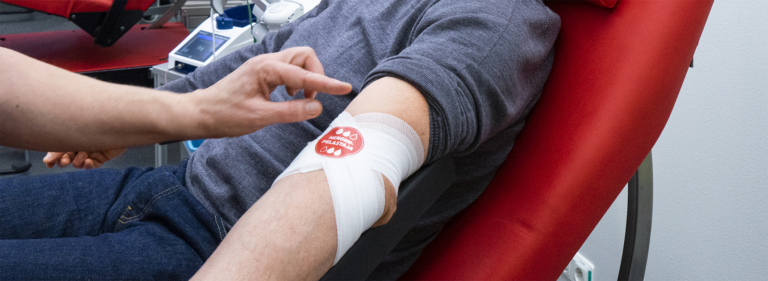
Blood donors are vital for patients
Blood donors only accounted for just over 3% of all people in the donating age range in Finland last year. Over the past ten years, the proportion has decreased by approximately a percentage point. It's…
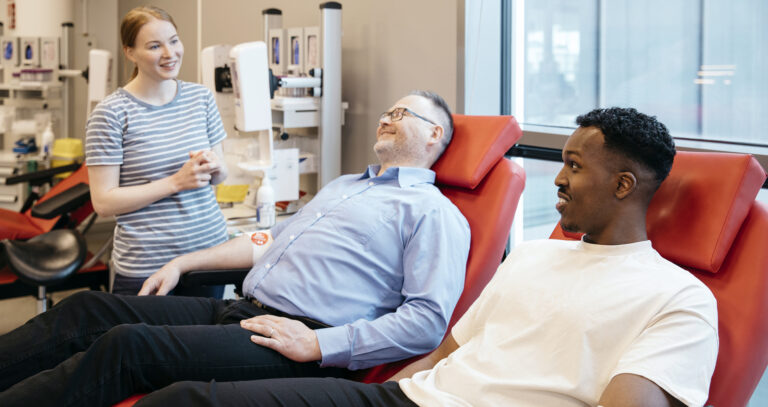
Check out the annual report 2023
How did we manage to take care of Finland's blood supply last year? In the Blood Service's new annual report 2023, you will find key figures and statistics, as well as reviews of the most important changes…

The application period for Blood Service research grants for 2024 is now open
The application period for 2024 grants from the Blood Service Research Fund has opened to researchers. The purpose of the grants is to support research related to current and future Blood Service products…